Chromatin Remodeling
| Epigenetics | Table of Contents |
It has long been known that the organization of eukaryotic DNA into chromatin is related to gene expression. For example, an early clue to this relationship was the observation that actively transcribed DNA is more susceptible to degradation by nucleases such as DNAse I suggesting that it is more "open" (less highly condensed by scaffolding proteins or into 30nm chromatin fiber) than non-transcribed DNA. In more recent years we have been learning a great deal more about how gene expression and chromatin structure are connected.
The basic idea of chromatin remodeling (also called nucleosome remodeling) was covered in the section on eukaryotic chromosome structure. What is discussed here is the relationship between the modification of histones and the changes in chromatin structure that result. These structural changes are what we refer to as chromatin remodeling.
Modifications associated with active transcription
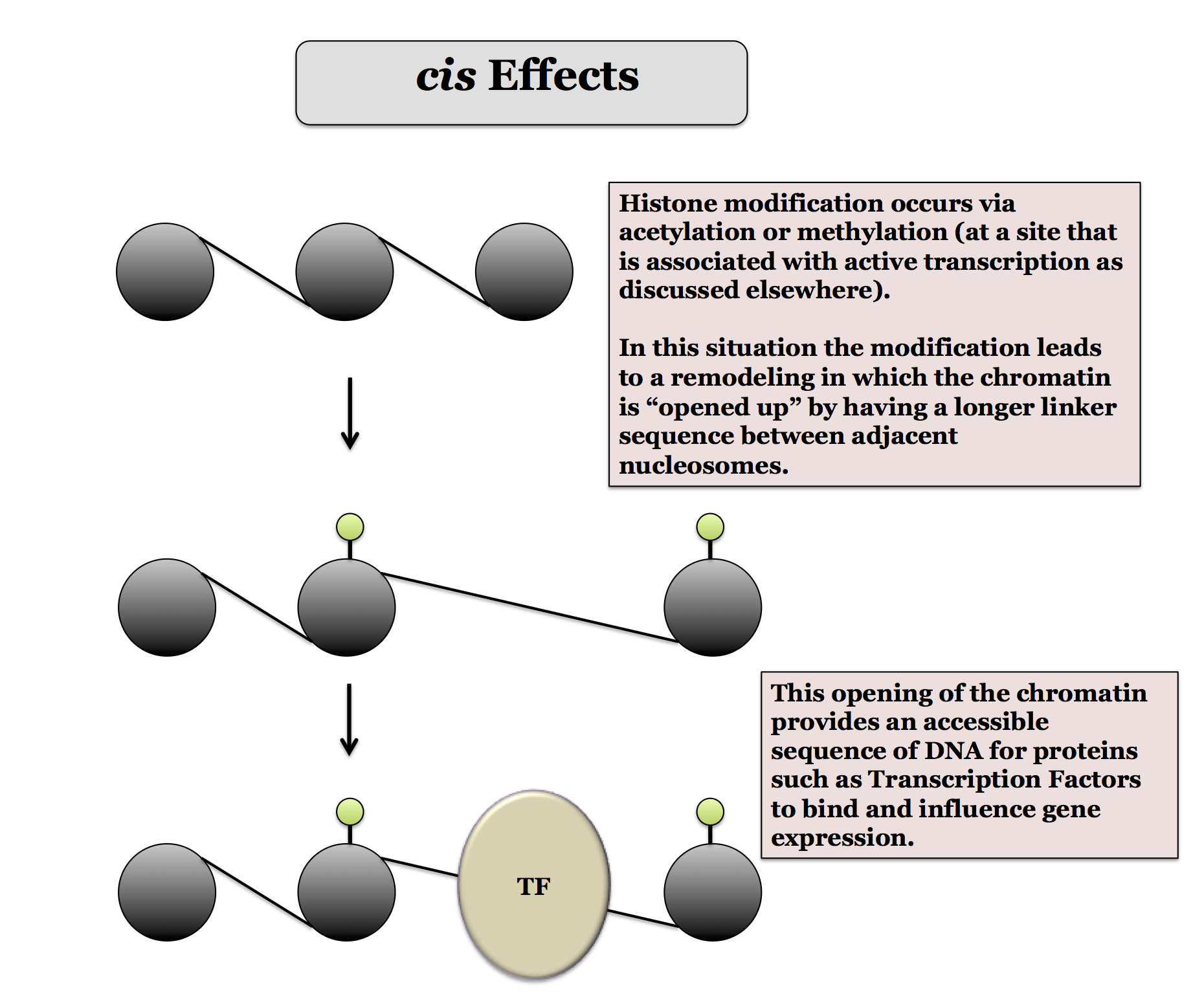
Remember that there are acetylations and some methylations that are associated with active transcription. The remodeling that these modifications lead to can be grouped into three types, cis-effects (changes in how nucleosomes are arranged, either by sliding nucleosomes or evicting them), trans-effects (changes brought about by a Reader biding to the modification) and histone exchange, in which histones in nearby nucleosomes are exchanged for other variants.
|
|
|
|
|
|
For each of the effects illustrated above the modification is caused by a writer protein such as an HAT (Histone Acetylase). The opposite effect can be achieved by an eraser protein, such as a HDAC (Histone Deacetlyase). So, for the cis-effects, an eraser could remove the modification and you would go from the open state back to the basic structure.
Therefore, gene expression can be down-regulated by reversing a modification. However, gene expression can also be down-regulated by causing changes that cause a remodeling that results in a structure that is inaccessible to proteins, thus causing very low levels of expression, perhaps even silencing. This is shown in the next section.
Modifications associated with gene silencing
Remember that there are specific methylations that are associated with inactive genes. These methylations lead to specific Reader proteins binding and inducing a remodeling in which the chromatin is condensed.
|
|
DNA Methylation and Chromatin Remodeling
|
|
| Epigenetics | Table of Contents |