Inversions
An inversion of a chromosome region is referred to as paracentric if the region does NOT include the centromere and pericentric if it does.
Inversions typically occur as a result of crossing over between different regions of the chromosome. If there are sections with inverted repeats that can align then crossing over will yield and inversion of the region between the repeats.
This is illustrated in this figure from the journal Nature.
Inversions frequently do not have any significant phenotypic effect. However, if either of the end points of the region is within a gene, the sequence of the gene will be disrupted in the inverted chromosome. This results in an amorphic allele which could have consequences for the phenotype.
Inversions and recombination
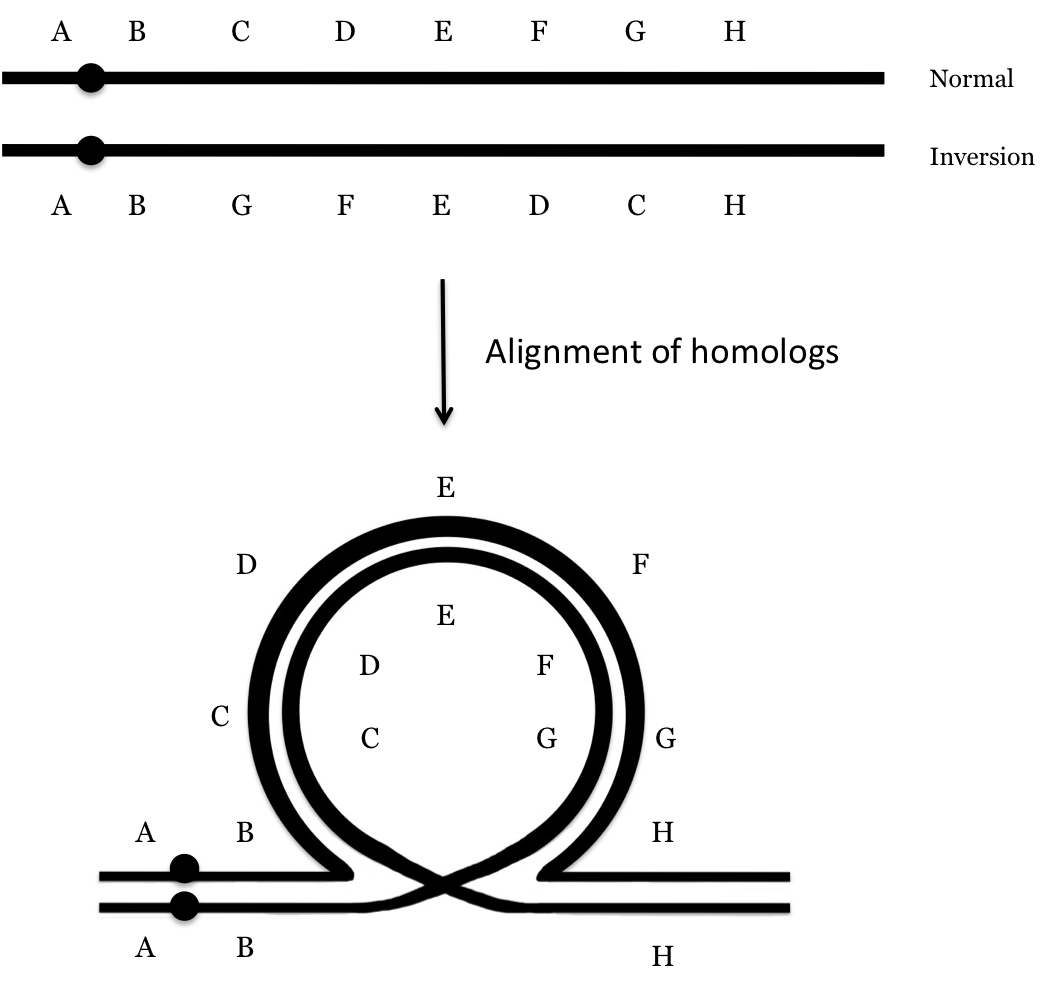
Although many chromosome inversions do not have a significant phenotypic effect, they can have an effect on genetic recombination. This will arise in individuals who are heterozygous for the inversion. At the first stage of meiosis in these individuals, the alignment of homologous chromosomes will require the formation of a loop in order to get all of the homologous sections lined up, as illustrated below for a paracentric inversion. In the diagram, letters represent some sort of genetic element along the chromosome, such as a gene or a band. For the sake of simplicity, only one chromatid of each homolog is drawn here but remember that when homologous chromosomes align, each has been replicated with the two sister chromatids being joined at the centromere.
|
|
If crossing over occurs within the inversion loop at this stage, no recombinant gametes will be formed. This is because the crossover in the loop results in the formation of aberrant chromosomes as seen in the following Figure. Only the chromatid involved in the crossing over are shown in the top portion in order to make it easier to see. The other two chromatid - those not involved in the crossover - simply segregate and are shown at the bottom.
|
|
To visualize how this happens, just mimic the processes of crossing over and meiosis. Break the chromatids that are involved in the cross over and rejoin them as the cell would. Then pull things apart by the centromeres as the spindle would during segregation - the two red arrows above connect a portion of each product to the piece of DNA in the looped DNA. Follow the chromatid that is attached to each centromere and pull it with that centromere. As shown in the diagrams, you should see that you get pieces of DNA that do not have the same content the original chromosomes. These are the chromatids that were involved in the crossing over.
As seen here in the case of a paracentric inversion the result of a cross over within the loop is one dicentric chromosome and an acentric fragment. The acentric fragment is eventually lost and degraded - no centromere = no good to a cell. The dicentric DNA will actually end up breaking at some point: the two centromeres will be segregating to opposite poles. Eventually that fragment will have to break during cell division - it cannot exist in both new cells. Where it breaks is random but regardless of where this occurs, neither cell will contain a full copy of the chromosome.
As a result of the aberrant chromosomes, the gametes carrying these products will not be viable. Therefore, no recombinant products are passed on, only chromosomes that were not involved in crossing over within the loop and these, of course, will have parental allele combinations for all genes within the inverted region. This is why inversions suppress recombination: they do not suppress crossing over, it is simply that the recombinant products are not viable.
Although not diagrammed here, crossing over within the loop formed by a pericentric inversion has the same effect of generating inviable products, and thus suppressing recombination, although some details differ.
Table of Contents