Complementation
Elsewhere we apply the concept of complementation when discussing the complementation test. Here we are just interested in the general concept of complementation as a form of gene interaction.
Complementation, by definition, is when two mutations produce a wild type phenotype when together in the same organism (or cell). Here we see how gene interaction lies at the heart of this process.
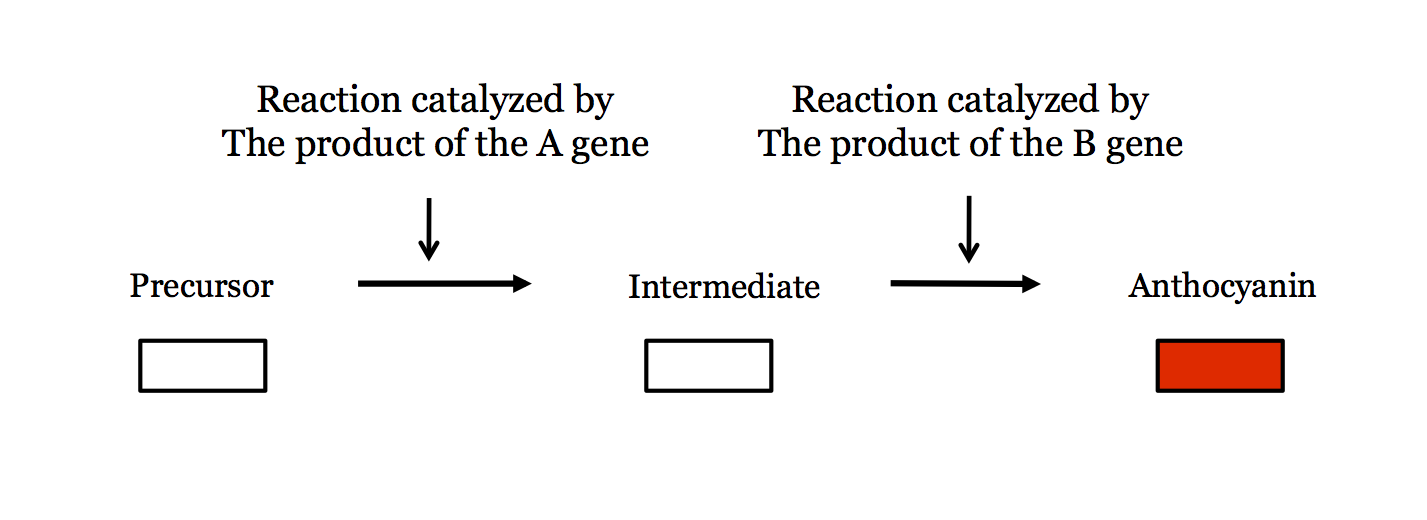
Consider the following hypothetical pathway for pigment synthesis in plants:
|
|
In our example there is a loss-of-function allele for the A gene and one for the B gene (Loss-of-function alleles are actually fairly common in biology so they are a good example.)
An individual who is homozygous for a will not produce functional enzyme and the first step of the pathway will not be catalyzed. As a result, no pigment will ever be produced and the plant will have white flowers. (Aa heterozygotes will express functional enzyme from the A copy and so they will catalyze the reaction: therefore, a is recessive).
Similarly, an individual who is homozygous for b will not produce functional enzyme and the second step of the pathway will not be catalyzed. As a result, the intermediate will build up but no pigment will ever be produced and the plant will have white flowers.
Knowing this, if you were to see a white-flowered plant you would have to ask: is it due to an aa genotype or to a bb genotype, or both? You cannot answer this just by observing the phenotype. You can, however, answer questions involving a comparison of two white-flowered plants. The most basic question would involve two different plants with white flowers: are they both homozygous for mutations in the same gene (that is both are aa or both are bb) or is one aaBB and the other AAbb?
What you should be able to see from the pathway is that if you cross an aaBB plant and an AAbb plant, both have white flowers but the progeny will be AaBb and have red flowers. This is complementation. The basic idea is that the although neither of the individual plants has all of the components (functional gene products) of a full pathway, the two plants between them have all of the components for a functional pathway - so they complement one another. From the diagram you can see how this complementation arises; the presence of A in the second plant complements the aa of the first and the B in the first plant complements the lack of a B allele in the second. As a result, the phenotype produced by the full pathway will show up in the progeny.
So, if the two white-flowered plants you crossed have mutations in different genes then they will complement each other and you will observe that in the progeny of the cross.
Conversely, if you cross two white flower plants that both happen to be aa, then the progeny will have white flowers, regardless of what the B genotypes are. So, the aa plants do not complement one another. Between them they do not have the components (gene products) of a functional pathway.
Complementation and dihybrid crosses.
Having considered the concept of complementation we can now look at what we would observe if you perform a dihybrid cross - AaBb x AaBb. As shown below you will observe a 9:7 phenotype ratio in the progeny. Without gene interaction you would get the classic 9:3:3:1 ratio but since the genotypes aaB_ (3/16), A_bb (3/16) and aabb (1/16) all have the same phenotype they are combined (7/16) in the phenotype ratio and so you get 9/16:7/16 or 9:7 as shown in this figure.
|
|