Dominant Epistasis
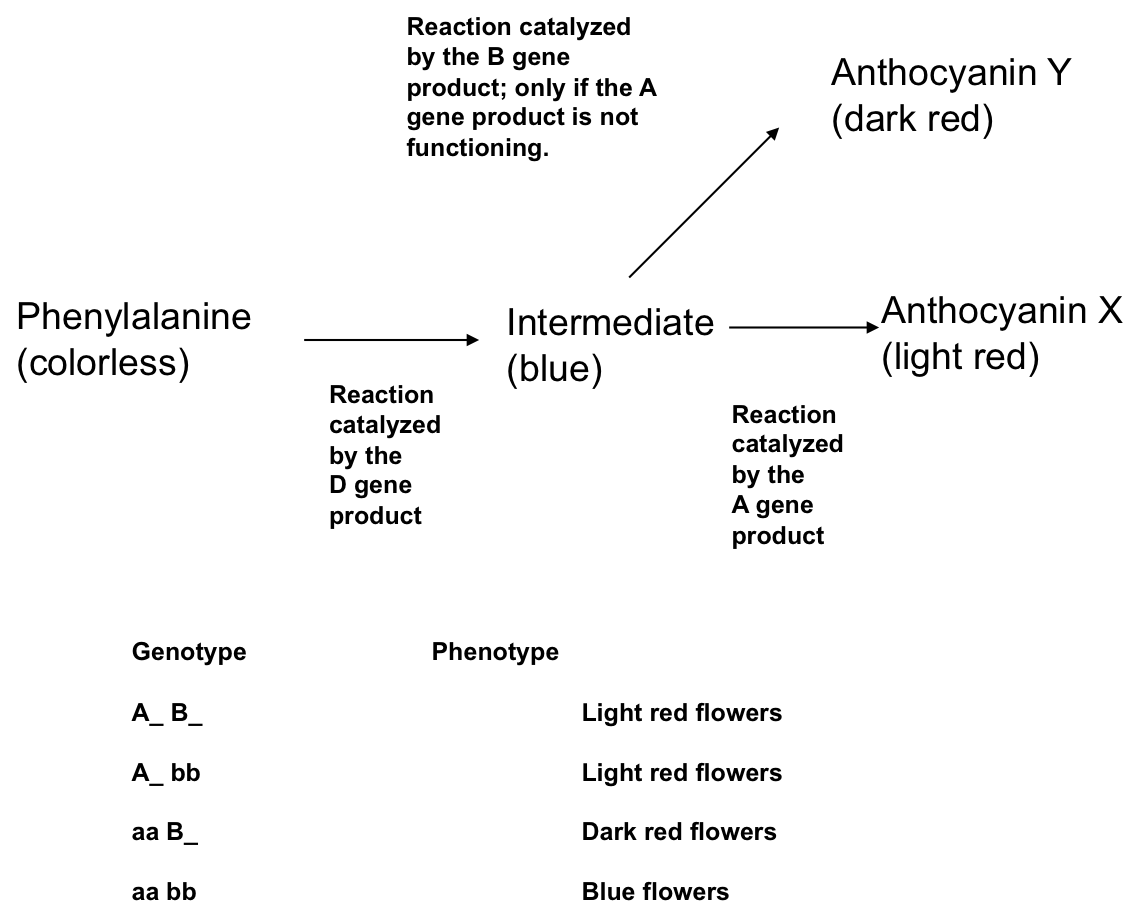
Dominant epistasis is a type of gene interaction in which one gene produces a phenotypic effect that masks the phenotypic effect of another gene. Unlike the case for recessive epistasis, however, it is the dominant trait for one gene that masks the effect of the second gene. For example, consider the pathway for plant pigments and alleles that are shown in the figure:
|
|
If this was the pathway that was responsible for pigment production, then the A gene would be epistatic to the B gene. As indicated in the figure, the B gene only catalyzes the reaction when the A gene is not acting (say in the case of an aa individual where a is a loss-of-function allele). It isn't particularly important why this might be the case since it is hypothetical, but one possibility would be that the enzyme coded by the B gene has a much lower reaction rate than the "A-coded"enzyme. If this were the case, then as soon as the substrate becomes available it will be converted by and A-coded enzyme before the B enzyme could act. In other words, the presence of A enzyme (in an individual that is AA or Aa) will be epistatic to B. Since the dominant A allele produces this epistatic effect, it is dominant epistasis. The relationship between genotype and phenotype for the A and B genes is shown in the figure. Of course, both of these depend on a functional D gene being present, the effect of which has been ignored here. (You might be able to see that the D gene would display recessive epistasis to both A and B.)
When a dihybrid cross is performed AaBb x AaBb you will observe a 12:3:1 phenotype ratio when there is dominant epistasis. Without gene interaction you would get the classic 9:3:3:1 ratio, but since A_B_ (9/16) and A_bb (3/16) have the same phenotype when A is epistatic to the B gene, they are combined (12/16) in the phenotype ratio.