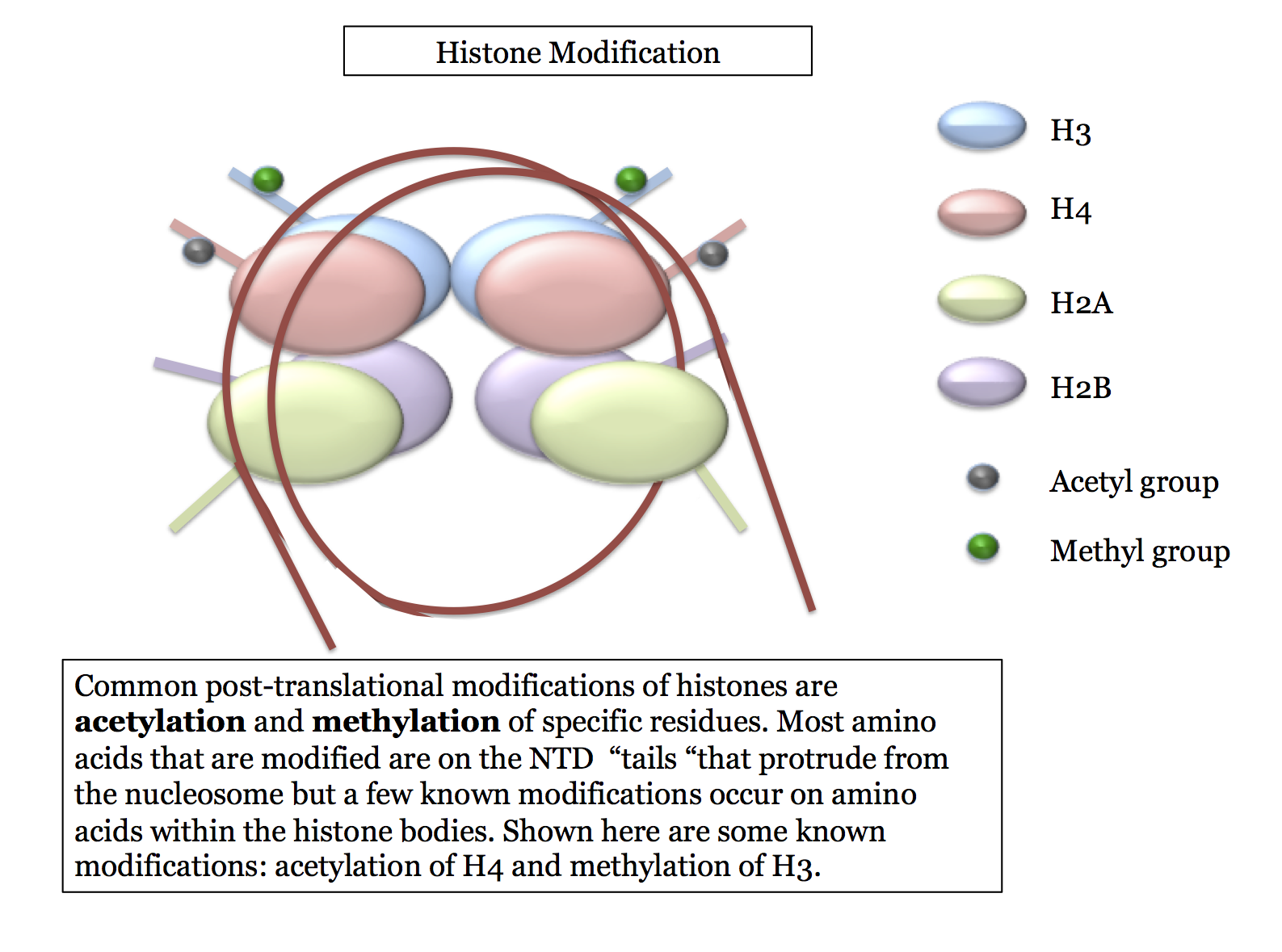
Histone Modification
| Eukaryotic Regulation | Table of Contents |
Histone modifications occur post-translationally (i.e. after the histone is translated) and are often changed as cellular conditions change: modifying particular histones is a common way for a cell to change the expression of genes coded on DNA near the nucleosome of that histone. The two most common modifications are:
There are other known modifications (such as phosphorylation and ubiquination) that we will not cover in our course. The ones we cover are those that seem to appear most frequently and which have been studied most thoroughly. Most, but not all, of these modifications occur at specific amino acids on the NTD - the N-Terminus Domains or "tails" that protrude from the nucleosome core. This is illustrated here.
|
|
Notation: Not every amino acid can be modified. Instead, as shown in the figures below there are specific amino acids that are modified; most commonly the modified amino acid is either a specific lysine (K), arginine (R) or serine (S) residue although other amino acids are sometimes modified. Furthermore, as the diagram indicates, most, but not all, modifications are found on histones H3 and H4. Histone modifications are designated so that both the histone and the amino acid modified are indicated as well as the specific modification. For example, the 16th amino acid in the sequence of H4 is a lysine and this lysine is a target for acetylation. When an H4 is acetylated it is designated by H4K16ac. In some cases, a residue can be modified by the addition of multiple groups: H3K4me3 indicates a trimethylated lysine in histone H3 - specifically the fourth amino acid of H3 (H3K4) which is a lysine. This notation is also given in the figures below.
Some histone modifications are associated with increased gene expression. Others are associated with decreased gene expression, or repression. The following two sections show you the acetylations and methylations associated with each.
Histone modifications associated with increased transcription rate
|
|
|
|
Histone modifications associated with decreased transcription rate
|
|
Effects
You should not try to memorize all of the specific modifications shown above. Rather, focus on the general concepts discussed next, which are based on the figures. The figures are just meant to give you an overall sense of the types of modifications and how many amino acids are targets of modifications.
Histone modifications affect the rate at which transcription initiation occurs for a gene that is near the nucleosome containing the histone(s) modified. This is turning out to be a very complex issue in genetics. The main difficult is that we have now found 60 different amino acids within histones that can be modified. This means that within any specific nucleosome there many possible combinations of modifications. And, since there are many nucleosomes near a gene the overall number of possible combinations across all of them is absolutely enormous. As a result it is proving to be a challenge to associate a specific modification with a specific effect on gene expression. Despite this, certain general effects are coming to light:
Histone methylation appears to influence the binding of non-histone proteins to the region. The binding of these proteins in turn affects various processes that play a role in transcription initiation. In some cases, histone methylation leads to the binding of proteins that then induce DNA methylation resulting in long-term silencing of the gene nearby.
More details about the relationship between histone modification and gene expression are given in the page on Chromatin Remodeling.
Proteins involved in histone modification
In the overview of epigenetics we discussed the types of proteins called writers and erasers. For acetylation and methylation there are specific writers and erasers. These are listed here; the functions are rather self-explanatory.
A video about acetylation:
| Eukaryotic Regulation | Table of Contents |