Mismatch Repair
| Repair and Recombination | Table of Contents |
Mismatch repair, as the name implies, repairs sites with normal bases but non-Watson Crick base pairing. (When there are abnormal bases at a site Base Excision Repair is utilized.) Most DNA polymerases have proofreading capability but even these will generate mis-pairings, and there are some polymerases involved in repair - particularly when the cell has undergone extensive damage - that lack proofreading or have very poor proofreading capability. Therefore, mis-pairings will exist and they need to be repaired.
What is discussed here is a specific post-replication repair mechanism in E. coli which is similar to systems found in other organisms. If there is a mismatch just following replication then it can be repaired, but the trick is to know which of the mis-paired bases is correct. If the cell can distinguish the template strand from the new strand then it can use the base in the template strand.
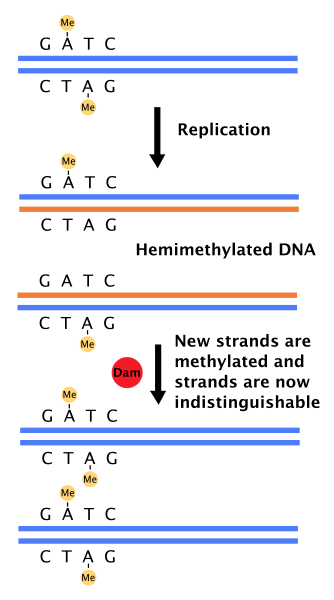
The enzyme Dam (DNA adenine methylase) is utilized for this. This enzyme methylates the Adenine at GATC sites throughout the genome (on both strands - it is self-complementary). It acts with a delay following replication so that the DNA is in a hemimethylated state for a period of time:
|
|
During the time that the DNA is hemimethylated, mismatch repair can use the methylated strand as the "correct" strand since the unmethylated DNA is obviously the newly synthesized strand. Repair then proceeds as diagrammed here:
|
|
A video covering mismatch repair.
| Repair and Recombination | Table of Contents |