Nucleosomes
| Euk. Chromosomes | Table of Contents |
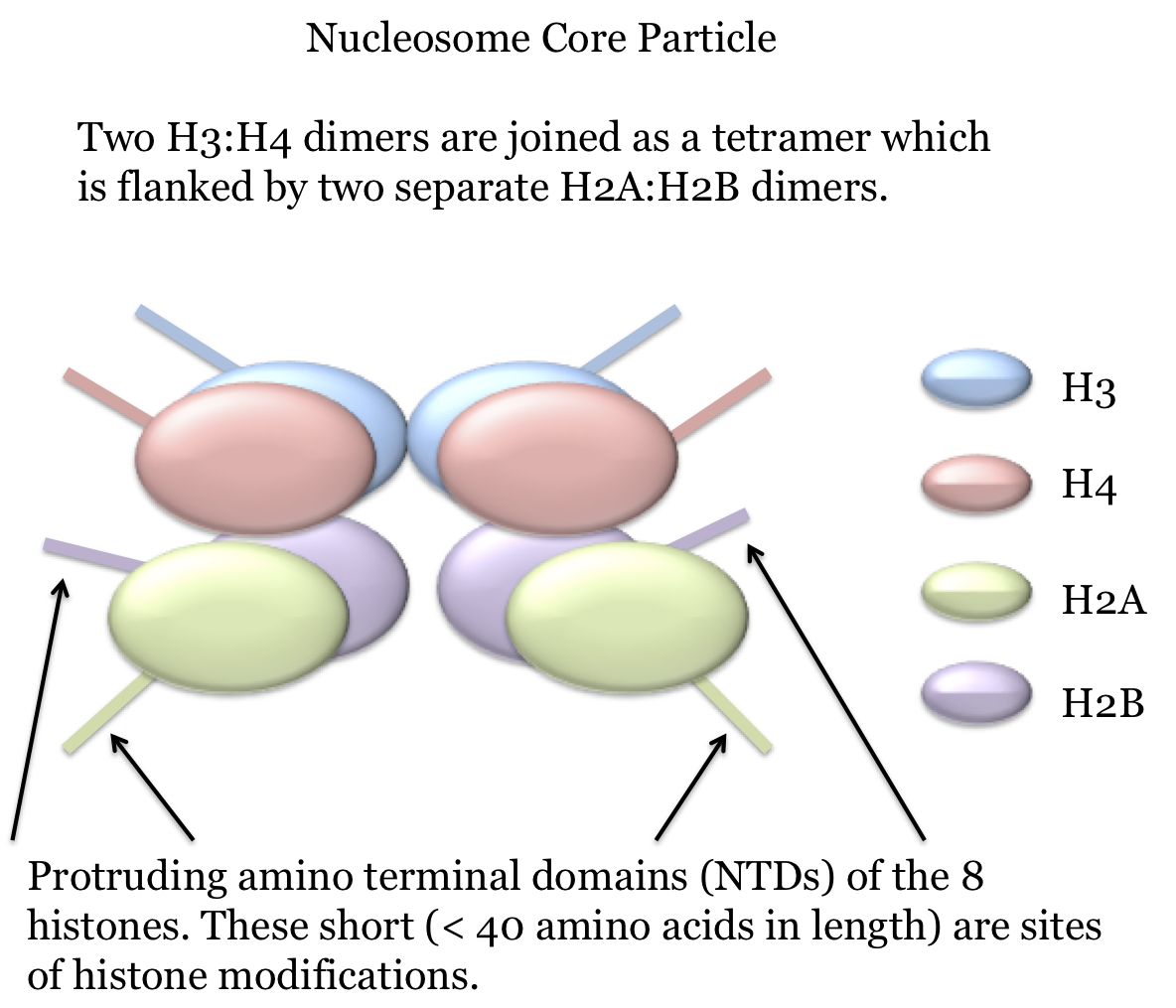
The basic structural unit of chromatin is the nucleosome. This is a structure composed of a core of 8 histone proteins - 2 copies each of histones H2A, H2B, H3 and H4 - around which 1 and 3/4 turns (about 146 nucleotides) of the double-stranded molecule is wound. Each histone has a stretch of roughly 40 amino acids at the amino terminus which extends like a tail out from the nucleosome. This tail - which is referred to as the NTD (N-Terminus Domain) - contains numerous basic residues and is where histone modifications, such as methyation and acetylation, occur. These modifications play important roles in a number of processes such as gene regulation and they will be discussed in various relevant sections.
Between adjacent nucleosomes there is a stretch of linker DNA that ranges in size from about 8 to just over 100 nucleotides depending on the species. A linker histone, H1, can bind to the nucleosome and form a structure sometimes referred to as a chromosome.
These structures are illustrated here.
|
|
|
|
|
|
Histone Variants: Although we describe the structure of a nucleosome in terms of an octamer composed of histones H2A, H2B, H3 and H4, eukaryotes actually have several forms of each histone, particularly H2A and H3 (H2B and H4 tend not to have different forms). The different forms of a particular histone are referred to as histone variants.
Organisms use different histone variants in different chromosome locations. The full consequences of this and the exact functions of each of the variants are not entirely understood but the general idea is that the variants affect nucleosome structure and can therefore affect chromatin structure. As a consequence of this they can affect processes such as DNA replication and gene expression.
Here are some examples of histone variants and where these variants are typically found. Although the actual function or purpose of each variant is not known, specific variants are found in specific chromosome locales, such as near a gene promoter. This suggests that the variant plays an important biological role related to that locale.
Histone H3Three common variants of histone H3 are called H3.1, H3.2 and H3.3. H3.1 and H3.2 are used in most nucleosomes, and are sometimes called the standard variants, but H3.3 is found in nucleosomes near transcription start sites of actively expressed genes. The incorporation of H3.3 is therefore thought to play a role in epigenetic regulation of gene expression. This type of histone exchange is covered in the section on histone modification, as aspect of epigenetic gene regulation.
Another variant of H3 is called CEN-P. As discussed below, this variant is abundant in nucleosomes in the centromere region.
Histone H2AAmong the variants of H2A are two called macroH2A and H2A.Z. The former is found in heterochromatin, including inactivated X chromosomes, while the H2A.Z form is found in higher concentrations in nucleosomes around the promoters of active genes.
Nucleosome Remodeling: This is also called chromatin remodeling, the terms are used in different places to refer to the same phenomenon. The organization of DNA into nucleosomes is dynamic: which stretches of a DNA molecule are wrapped around a core of histones can change. This is an important feature of chromatin structure because the interaction between DNA and histones can present problems for the binding of other proteins to the DNA, something necessary of many critical processes such as transcription and replication. When a stretch of DNA is in tight interaction with the core histones it is less accessible than if it is free, such as in linker DNA. By changing the arrangement of nucleosomes the cell can make a specific sequence of DNA accessible to proteins, an important aspect of eukaryotic genetics.
The following figures show the three major ways in which chromatin remodeling can occur. The first two involve a change in what DNA is wrapped around the core, thus making a specific sequence accessible for protein binding. This can occur by sliding a nucleosome along the DNA or by removing (evicting) a nucleosome. These changes are performed by proteins called Chromatin Remodeling Factors (or Nucleosome Remodeling Factors). One well-known example is the SWI/SNF complex.
|
|
|
|
Chromatin remodeling also includes histone exchange. This is a process by which the histone variants of a specific nucleosome are exchanged for different variants as illustrated here. This does not change DNA accessibility like the sliding and eviction processes shown above but the resulting change in nucleosome structure can be important for various processes or general chromosome structure. (These are not discussed here but may be mentioned in different pages covering molecular processes.)
|
|
Chromatin remodeling is most frequently mentioned in association with gene regulation. This is because much of the remodeling occurs around gene regions and affects transcription levels due to the accessibility or inaccessibility of DNA to Transcription Factors. Also, it is the process in which chromatin remodeling has been studied most extensively. Therefore, mechanistic details about remodeling will be covered in the section on epigenetic and gene regulation.
| Euk. Chromosomes | Table of Contents |