Applying Recombination Frequency
Once you have calculated Recombination Frequency (RF) for any two genes (or get it from a linkage map) you can apply this knowledge to make predictions about genetic cross results. As long as there is no chromosome mutation or rearrangement, the two genes will always display the same frequency of recombination.
As you know, the main problem with linkage is that the genes do not assort independently. Therefore, it is not a simple matter to determine the frequency with which progeny phenotypes will appear.
Test Crosses
We will start with test crosses since they are fairly simple. Remember that RF is defined from a test cross: the RF between two genes is defined as the number of recombinant progeny that are observed in a test cross.
So, if the test cross involves A B/a b (two genes in cis configuration) then RF is calculated by the number of progeny that inherit a recombinant gamete ([A,b] or [a,B]) divided by the total number of progeny. Make sure you review that if you are not comfortable with the calculation.
Given this, we can very easily turn it around. If, instead, you KNOW the RF then you can apply it as follows: if two genes have an RF value of 10%, then, by the definition, when you do a test cross, you expect 10% of the progeny to inherit a recombinant gamete (and 90% a parental gamete).
So, from the cross A B/a b x aabb we expect the following progeny proportions:
Since for every [A, B] gamete there is an [a, b] gamete that resulted from segregation, and for every [A, b] gamete there is an [a, B] gamete as a result of segregation, we expect the [A, B]:[a, b] ratio to be 1:1 and the [A, b]:[a, B] ratio to be 1:1.
The result is that we expect to observe the following from the cross:
If the cross had involved a trans configuration heterozygote then:
For a cis configuration test cross, this can easily be generalized to:
For a trans configuration cross we would swap the Parental and Recombinant phenotypes accordingly.
A very important thing to understand is that these values are also telling you the expected gamete frequencies for the heterozygote. Since the progeny phenotype from a test cross is determined by the gamete inherited from the heterozygote parent, progeny frequencies are the same as gamete frequencies.
Dihybrid Crosses
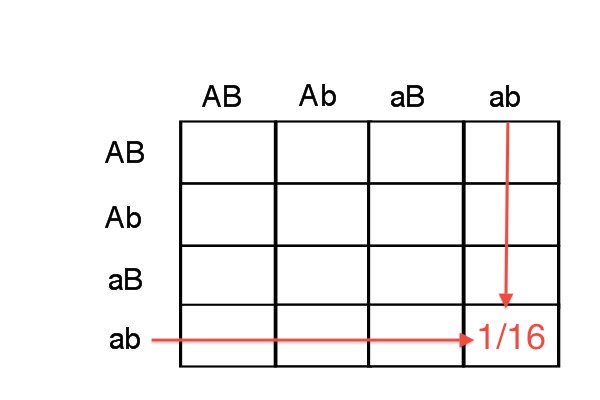
Make sure that you follow the concept above about the expected gamete frequencies from a heterozygote (the last sentence). When you do a dihybrid cross it is a fairly simple matter to calculate the probability of inheriting two particular gametes. You have done this before using a Punnett Square:
|
|
The probability of getting aabb is the probability of getting an [a, b] gamete AND an [a, b] gamete which is 1/4 * 1/4 or 1/16.
What is important here is to see where the gamete frequency of 1/4 came from. This is a result of Independent Assortment. From a heterozygote, 1/2 the gametes will carry [A] and 1/2 the gametes will carry [a]. Similarly, 1/2 will carry [B] and 1/2 will carry [b]. If the genes segregate independently, the Pr(a AND b) = Pr(a) * Pr(b) = 1/2 * 1/2 = 1/4.
This is where linkage creates a problem; the genes do not segregate independently so we cannot apply the AND rule when calculating gamete frequencies. We can, however, still apply the AND rule to zygote frequencies since the two gametes are independent of one another. Therefore, we can still apply the Punnett Square approach as long as we know gamete frequencies.
You may see where this is going. As we saw above, we can use RF to calculate gamete frequencies from a heterozygote. Therefore, we can use this approach to find gamete frequencies and then apply these to a Punnett Square. So, using the information about RF we can fill in gamete frequencies on a Punnett Square. For example, if we cross two cis configuration individuals, then the gamete frequencies are as shown:
|
|
We simply alter these if either, or both, is in trans configuration (which just changes which gametes are parental and which are parental).
We can now use the AND rule to calculate any square (or all of them if we wanted). For example Pr(aabb) is still Pr([ab] AND [ab]) so we multiply as with independent genes, it's just that we aren't multiplying 1/4 and 1/4 but instead:
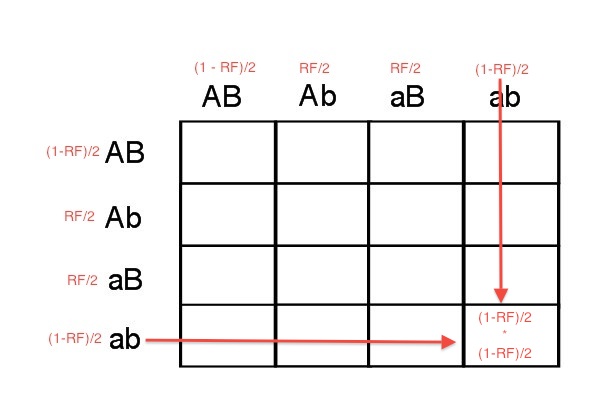
From the cross of cis individuals, the probability of an aabb progeny is (1-RF)/2 * (1/RF)/2.
As an example, if the two genes are 40 cM apart (40%RF), we expect 4% of the progeny to be aabb:
|
|
As we have done before with Punnett Squares, if we are interested in a particular genotype or phenotype then we calculate each appropriate square and add them together.